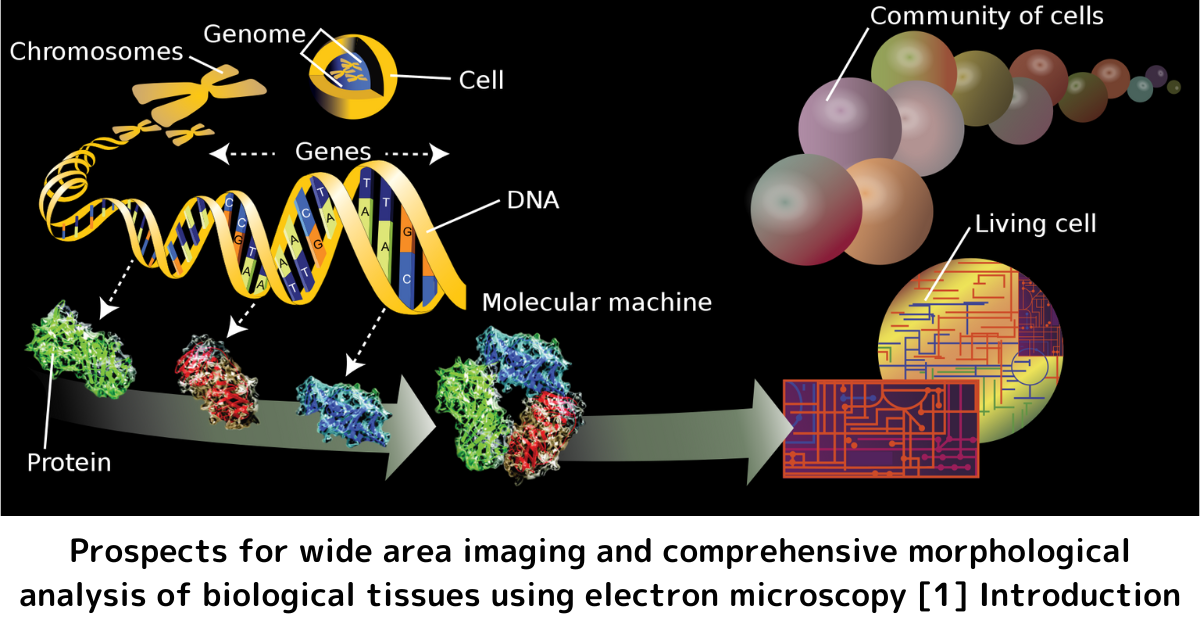
The above illustration is reprinted from : Omics - Wikipedia
Introduction
Biological tissues contain countless amounts of extremely fine and complex morphological information. In order to understand such complex morphological information, visualization techniques such as microscopy, imaging techniques and tissue staining methods have been developed so far.
In particular, electron microscopy, the highest resolution imaging method, is used to observe the ultra-fine morphological structures in biological tissues. The morphological information obtained by electron microscopy includes nano-scale information on the tissues and cells, such as the shapes of cell nuclei, the state of chromatin DNA, the morphology of intracellular organelles such as mitochondria, Golgi apparatus, and endoplasmic reticulum (ER), as well as the neural networks and synaptic structure between neurons in the central nervous system of brain.
Biological morphome
The term "Morphome" (also "biological morphome") is used to refer to the aggregate of morphological characteristics in biological tissues and cells. The morphome such as the morphological information of tissues and cells is expressed as the sum of molecular dynamics such as DNA information (genome) and RNA information (transcriptome), and it is also regarded as morphological phenotype information. Most of the morphological data are image data, which are seemingly very different from the sequence data and numerical data mainly used in the conventional omics data (genomics, transcriptomics and metabolomics) of molecular biology.
Morphomics
Morphomics, a research field to handle a large-scale of morphological data, is now emerging through the application of the latest imaging and information processing technologies. This means that the quantitative information on morphological images is treated as new omics information by combining comprehensive morphological information and bioimage informatics.
In this article, as one of the imaging techniques for comprehensively acquiring morphological information of biological tissues, I introduce a scanning electron microscope (SEM) for the wide-area imaging of tissue sections, combined with an array tomography method for producing continuous ultra-thin sections. For bioimaging and informatics research, it is also important to try to identify various morphological information from the large-scale image information "image big data", and to analyze the shape and 3D volume of cells quantitatively. As one of the technologies contributing to the realization of such image processing and analysis, I outline deep learning (DL) and its applications.